Bacteria have remarkable capacity of adaption to various ecosystems. Our team interests in the questions about how magnetotactic bacteria develop their swimming apparatus to exploit the geomagnetic field and what are the mechanisms used by deep-sea bacteria to draw energy from hydrothermal vent light. The model organisms include unicellular magneto-spirillum, magneto-coccus and multicellular magnetotactic magnetoglobules collected from the Mediterranean Sea and China Sea, and both gram-positive and gram-negative bacteria collected and isolated from hydrothermal vent fields in Indian Ocean. These studies are carries out using genomic, molecular, biochemistry, structure, metagenomic, microscopic and biophysical approaches through national and international collaborations.
Motility is the ability of an organism to move by itself. The motility of a cell may enable it to move toward a desired environment, or to flee from a harmful environment. Bacterial cells use structures called flagella to enable this movement. Bacteria presenting flagella at each pole are said amphitrichous. While motility models for peritrichous or monotrichous bacteria are well established, little is known on motility of amphitrichous bacteria. Moreover, some of these species glycosylate their flagellins and this process is crucial for the formation of a functional flagella filament and thus for cell motility. We investigate the flagella glycosylation and the motility of the amphitrichous magnetotactic Magnetospirillum magneticum AMB-1. The biochemical analysis of the flagellin of AMB-1 revealed that this protein was glycosylated. By deletion of a gene (called maf) located near the only functional flagellin gene, we demonstrated that its product is a GT involved in the transfer of a sugar onto the flagellin and, more importantly, that this glycosylation is essential for the assembly of flagellar filaments. We also determined the 3-D structure of Maf (Fig1).
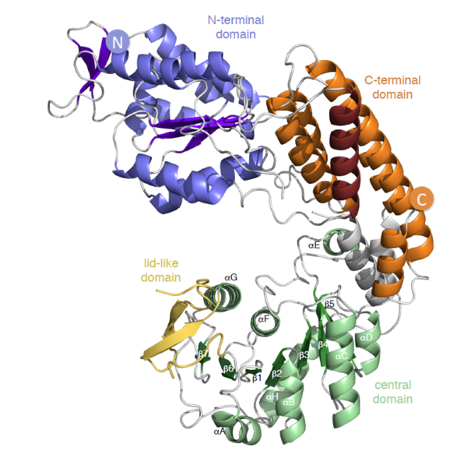
Legend Fig1 : Maf structure colored by domains: the N-Ter is represented in blue, the central domain in green and the C-Ter in orange. The boundaries of the disordered region corresponding to the cover domain of the GT42 family are in yellow. The L92-S108 helix participating in the C-terminal domain is in ruby, and the helices of the inter-domain links are in gray. The secondary structural elements of the central domain, β1 – β7 and αA – αH are marked (PDB: 5MU5).
In a previous work we described the three motility behaviors of an amphitrichous bacterium: runs, tumbles, and reversals. We identified two characteristic patterns corresponding to flagella rotating in opposite directions: “parachutes” (labeled P in Fig. 2) corresponding to a clockwise rotation of the flagellum (CW) and “tuft” positions (labeled T in Fig. 2) to a counterclockwise rotation (CCW).
Legend Fig2 : Schematic representation of the 3 motility behaviors of AMB-1 cells. The two patterns: parachute (P) or tuft (T) are given for each flagellum for each behavior. AMB-1 cell representation: green the cell body and in pink the flagella.
We concluded that an amphitrichous bacterium is able to operate its two flagella differently at the poles and needs to coordinate them precisely when it changes direction. We are now exploring the molecular mechanism at the origin of this asymmetric behavior.
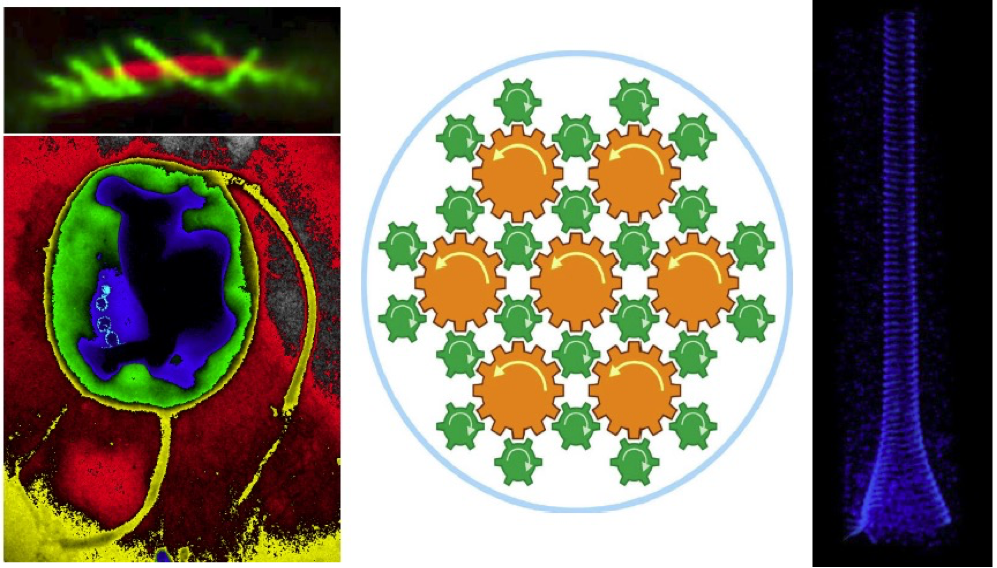
Light serves as signal and energy for marine bacteria. Magnetoglobules are multicellular gram-negative bacteria dwelling in marine shallow water sediments. Approximately 60 phylogenetically identical cells assemble into a single layer ellipsoidal entity with thousands of flagella at the out surface (Fig. 1). They align and swim along the magnetic field lines, a behavior that is referred to as magnetotaxis, to move to favorable oxic-anoxic interface in chemically stratified sediments. In response to illumination of light magnetoglobules reverse suddenly their swimming direction with acceleration. We have sequenced genomes of 5 species of ellipsoidal magnetoglobules collected from the Mediterranean Sea and China Sea, and identified genes involved in circadian rhythm. The photosensitive magnetotaxis might be selective strategies for magnetoglobules to move to favorable places in accordance with the daily nutrient displacement in sediments. The other model organisms studied in our team are deep-sea phototrophic bacteria. Deep-sea hydrothermal vents are known as chemosynthetic ecosystems. However, high temperature vents emit light that hypothetically can drive photosynthesis in this habitat. To contribute to a comprehensive understanding of phototrophic microbial diversity in this ecosystem we collected samples from hydrothermal vent on the Southwest Indian Ridge. Metagenomic analysis identified populations of known potential chlorophototrophs and retinalophototrophs. Complete chlorophyll biosynthetic pathways were identified in all samples. By contrast, proteorhodopsins were only found in active beehive smoker diffusers. In addition, we have isolated vent bacteria and succeeded to boost vent microbial growth by, for the first time, solely infrared illumination under in-situ simulation conditions. We are investigating the phototrophic mechanism and evaluating the contribution of the geothermal light to the vent ecosystems that is independent on solar energy.
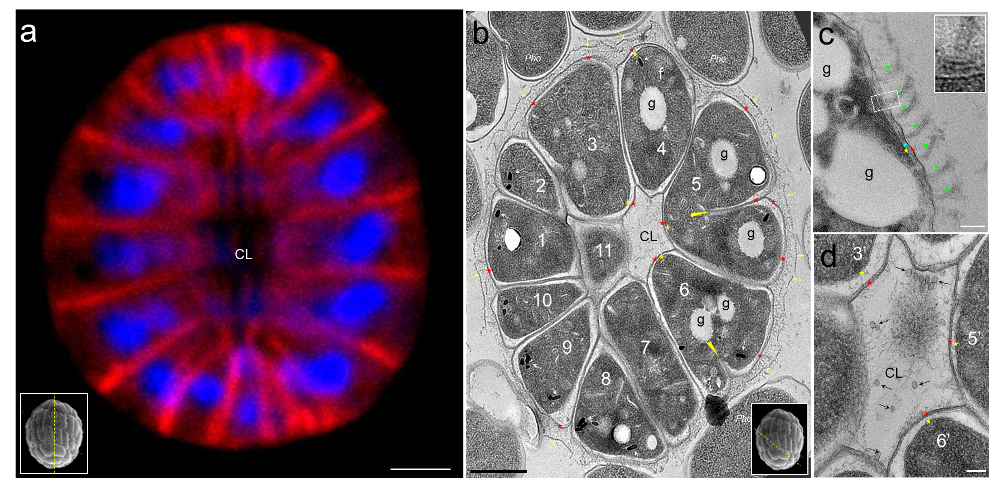
Legend Fig 1 : Ultrastructure of ellipsoidal magnetoglobules. (a) shows a center stack or reconstitution of laser confocal images of ellipsoidal magnetoglobules; the membrane is stained by FM4-64 (red) and DNA by DAPI (blue). (b to d) are TEM-bright field (BF) mode micrographs of ultrathin sections of HPF/FS-fixed ellipsoidal magnetoglobules. (b) shows cells surrounding a core lumen (CL). The yellow arrows visualize the dashed line on the surface of the magnetoglobule and the red and yellow asterisks indicate the outer and the inner membrane, respectively. White arrows point to the magnetosomes and the “f” is a fence-like structure. The yellow triangles indicate the extremity of the growing septa in cells 5 and 6. (c) shows flagella (green triangles) and a magnified image (inset) with likely the hook and basal body of a flagellum. (d) is a magnified view of the core lumen (CL) with vesicles (black arrows) and filaments. “g” are Nil red-stainable lipid granules. Scale bars: 1 µm in (a) and (b), 100 nm in c and d images. The insets in (a) and (b) illustrate the estimated cut direction of the corresponding sections.
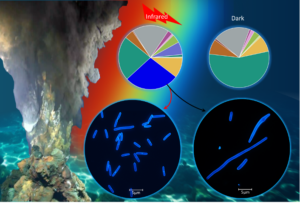
Legend Fig 2 : The infrared radiation from hydrothermal vents impacts the composition of the microbial population, promoting bacterial growth by triggering cell division as an adaptive strategy for living in deep-sea habitats. (Dai, J., Li, X.-G, et al. (2024). Illuminating a bacterial adaptation mechanism: Infrared-driven cell division in deep-sea hydrothermal vent environments. The Innovation Geoscience, 100050. 10.59717/j.xinn-geo.2024.100050.)
Group leader / Research director (DR-CNRS)
Researcher (CDD-AMU)
Assistant professor (CDD-AMU)
Researcher (CDD-AMU)
PhD student (PhD-AMU)
PhD student (PhD-AMU)



