Science – New Method for Analyzing Microbiome Interactions
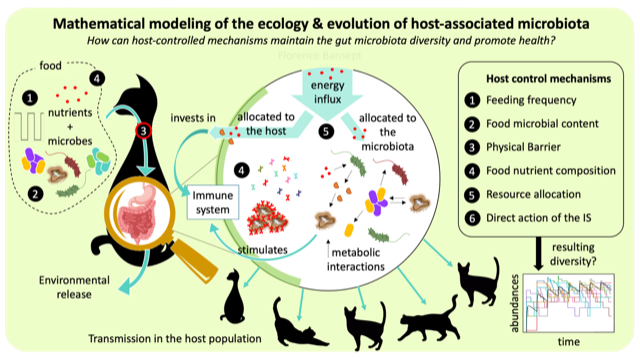
New Method for Analyzing Microbiome Interactions From R. Zapién-Campos, F. Bansept, A. Traulsen Researchers Develop Stochastic Approach for More Accurate Models of Microbial Communities. Microbial